Links
Databases
 OPM Orientations of Proteins in Membranes (OPM) database
OPM Orientations of Proteins in Membranes (OPM) database
PDBTM PDBTM, Protein Data Bank of Transmembrane Proteins
 Membrane Proteins of Known 3D Structure, a current list of membrane protein structures determined by x-ray and electron diffraction with links to the Protein Data Bank and other useful sites.
Membrane Proteins of Known 3D Structure, a current list of membrane protein structures determined by x-ray and electron diffraction with links to the Protein Data Bank and other useful sites.
Tools
DOWSER program The DOWSER program surveys a protein molecule's structure to locate internal cavities and assess the hydrophilicity of these cavities in terms of the energy of interaction of a water molecule with the surrounding atoms.
Our web services
 SuperLooper provides an online interface for the automatic, quick and interactive search and placement of loops in proteins.
SuperLooper provides an online interface for the automatic, quick and interactive search and placement of loops in proteins.
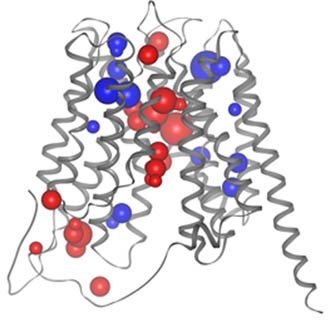 Voronoia is a program suite to analyze and visualize the atomic packing of protein structures.
Voronoia is a program suite to analyze and visualize the atomic packing of protein structures.
 Voronoia for RNA, a database storing data on atomic packing densities of RNA structures and complexes.
Voronoia for RNA, a database storing data on atomic packing densities of RNA structures and complexes.
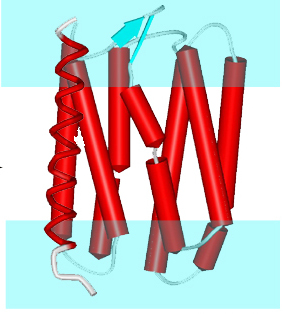 Rhythm is a server to predict the orientation of transmembrane helices in channels and membrane-coils.
Rhythm is a server to predict the orientation of transmembrane helices in channels and membrane-coils.
 Version: 2014-05-21
Version: 2014-05-21