Interface of β2 Adrenergic Receptor in Complex with Empty Gs-Protein¶
[1]:
import mdtraj as md
import mdciao
Download example data and load into the namespace¶
[2]:
import numpy as np
import os
if not os.path.exists("mdciao_example"):
mdciao.examples.fetch_example_data("b2ar@Gs")
traj = md.load("mdciao_example/traj.xtc",top="mdciao_example/prot.pdb")
Unzipping to 'mdciao_example'
Create consensus labeler objects¶
[3]:
GPCR = mdciao.nomenclature.LabelerGPCR("adrb2_human")
CGN = mdciao.nomenclature.LabelerCGN("3SN6")
No local file ./adrb2_human.xlsx found, checking online in
https://gpcrdb.org/services/residues/extended/adrb2_human ...done!
Please cite the following reference to the GPCRdb:
* Kooistra et al, (2021) GPCRdb in 2021: Integrating GPCR sequence, structure and function
Nucleic Acids Research 49, D335--D343
https://doi.org/10.1093/nar/gkaa1080
For more information, call mdciao.nomenclature.references()
done without 404, continuing.
Using CGN-nomenclature, please cite
* Flock et al, (2015) Universal allosteric mechanism for G$\alpha$ activation by GPCRs
Nature 2015 524:7564 524, 173--179
https://doi.org/10.1038/nature14663
No local file ./CGN_3SN6.txt found, checking online in
https://www.mrc-lmb.cam.ac.uk/CGN/lookup_results/3SN6.txt ...done without 404, continuing.
No local PDB file for 3SN6 found in directory '.', checking online in
https://files.rcsb.org/download/3SN6.pdb ...found! Continuing normally
Guess molecular fragments¶
This would be done anyway by the mdciao.cli.interface call in the cell below, here we do it have the fragments defined in the namespace
[4]:
fragments = mdciao.fragments.get_fragments(traj.top);
fragment_names = ["Galpha", "Gbeta", "Ggamma","B2AR","P0G"]
Auto-detected fragments with method 'lig_resSeq+'
fragment 0 with 354 AAs LEU4 ( 0) - LEU394 (353 ) (0) resSeq jumps
fragment 1 with 340 AAs GLN1 ( 354) - ASN340 (693 ) (1)
fragment 2 with 66 AAs ALA2 ( 694) - PHE67 (759 ) (2)
fragment 3 with 283 AAs GLU30 ( 760) - LEU340 (1042 ) (3) resSeq jumps
fragment 4 with 1 AAs P0G395 ( 1043) - P0G395 (1043 ) (4)
Compute G\(\alpha\)-B2AR interface¶
Using the above fragment definitions
[5]:
intf = mdciao.cli.interface(traj,
title="3SN6 beta2AR-Galpha interface",
fragments=fragments, fragment_names = fragment_names,
frag_idxs_group_1=[0],
frag_idxs_group_2=[3],
GPCR_uniprot=GPCR, CGN_PDB=CGN,
accept_guess=True, no_disk=True, figures=False)
Will compute contact frequencies for trajectories:
<mdtraj.Trajectory with 280 frames, 8384 atoms, 1044 residues, and unitcells>
with a stride of 1 frames
Using method 'user input by residue array or range' these fragments were found
fragment 0 with 354 AAs LEU4 ( 0) - LEU394 (353 ) (0) resSeq jumps
fragment 1 with 340 AAs GLN1 ( 354) - ASN340 (693 ) (1)
fragment 2 with 66 AAs ALA2 ( 694) - PHE67 (759 ) (2)
fragment 3 with 283 AAs GLU30 ( 760) - LEU340 (1042 ) (3) resSeq jumps
fragment 4 with 1 AAs P0G395 ( 1043) - P0G395 (1043 ) (4)
GPCR-labels align best with fragments: [3] (first-last: GLU30-LEU340).
These are the GPCR fragments mapped onto your topology:
TM1 with 32 AAs GLU30@1.29x29 ( 760) - PHE61@1.60x60 (791 ) (TM1)
ICL1 with 4 AAs GLU62@12.48x48 ( 792) - GLN65@12.51x51 (795 ) (ICL1)
TM2 with 32 AAs THR66@2.37x37 ( 796) - LYS97@2.68x67 (827 ) (TM2)
ECL1 with 4 AAs MET98@23.49x49 ( 828) - PHE101@23.52x52 (831 ) (ECL1)
TM3 with 36 AAs GLY102@3.21x21 ( 832) - SER137@3.56x56 (867 ) (TM3)
ICL2 with 8 AAs PRO138@34.50x50 ( 868) - LEU145@34.57x57 (875 ) (ICL2)
TM4 with 27 AAs THR146@4.38x38 ( 876) - HIS172@4.64x64 (902 ) (TM4)
ECL2 with 20 AAs TRP173 ( 903) - THR195 (922 ) (ECL2) resSeq jumps
TM5 with 42 AAs ASN196@5.35x36 ( 923) - GLU237@5.76x76 (964 ) (TM5)
ICL3 with 2 AAs GLY238 ( 965) - ARG239 (966 ) (ICL3)
TM6 with 35 AAs CYS265@6.27x27 ( 967) - GLN299@6.61x61 (1001 ) (TM6)
ECL3 with 4 AAs ASP300 ( 1002) - ILE303 (1005 ) (ECL3)
TM7 with 25 AAs ARG304@7.31x30 ( 1006) - ARG328@7.55x55 (1030 ) (TM7)
H8 with 12 AAs SER329@8.47x47 ( 1031) - LEU340@8.58x58 (1042 ) (H8)
CGN-labels align best with fragments: [0] (first-last: LEU4-LEU394).
These are the CGN fragments mapped onto your topology:
G.HN with 28 AAs THR9@G.HN.26 ( 5) - VAL36@G.HN.53 (32 ) (G.HN)
G.hns1 with 3 AAs TYR37@G.hns1.1 ( 33) - ALA39@G.hns1.3 (35 ) (G.hns1)
G.S1 with 7 AAs THR40@G.S1.1 ( 36) - LEU46@G.S1.7 (42 ) (G.S1)
G.s1h1 with 6 AAs GLY47@G.s1h1.1 ( 43) - GLY52@G.s1h1.6 (48 ) (G.s1h1)
G.H1 with 7 AAs LYS53@G.H1.1 ( 49) - GLN59@G.H1.7 (55 ) (G.H1)
H.HA with 26 AAs LYS88@H.HA.4 ( 56) - LEU113@H.HA.29 (81 ) (H.HA)
H.hahb with 9 AAs VAL114@H.hahb.1 ( 82) - PRO122@H.hahb.9 (90 ) (H.hahb)
H.HB with 14 AAs GLU123@H.HB.1 ( 91) - ASN136@H.HB.14 (104 ) (H.HB)
H.hbhc with 7 AAs VAL137@H.hbhc.1 ( 105) - PRO143@H.hbhc.15 (111 ) (H.hbhc)
H.HC with 12 AAs PRO144@H.HC.1 ( 112) - GLU155@H.HC.12 (123 ) (H.HC)
H.hchd with 1 AAs ASP156@H.hchd.1 ( 124) - ASP156@H.hchd.1 (124 ) (H.hchd)
H.HD with 12 AAs GLU157@H.HD.1 ( 125) - GLU168@H.HD.12 (136 ) (H.HD)
H.hdhe with 5 AAs TYR169@H.hdhe.1 ( 137) - ASP173@H.hdhe.5 (141 ) (H.hdhe)
H.HE with 13 AAs CYS174@H.HE.1 ( 142) - LYS186@H.HE.13 (154 ) (H.HE)
H.hehf with 7 AAs GLN187@H.hehf.1 ( 155) - SER193@H.hehf.7 (161 ) (H.hehf)
H.HF with 6 AAs ASP194@H.HF.1 ( 162) - ARG199@H.HF.6 (167 ) (H.HF)
G.hfs2 with 5 AAs CYS200@G.hfs2.1 ( 168) - GLY206@G.hfs2.7 (172 ) (G.hfs2) resSeq jumps
G.S2 with 8 AAs ILE207@G.S2.1 ( 173) - VAL214@G.S2.8 (180 ) (G.S2)
G.s2s3 with 2 AAs ASP215@G.s2s3.1 ( 181) - LYS216@G.s2s3.2 (182 ) (G.s2s3)
G.S3 with 8 AAs VAL217@G.S3.1 ( 183) - VAL224@G.S3.8 (190 ) (G.S3)
G.s3h2 with 3 AAs GLY225@G.s3h2.1 ( 191) - GLN227@G.s3h2.3 (193 ) (G.s3h2)
G.H2 with 10 AAs ARG228@G.H2.1 ( 194) - CYS237@G.H2.10 (203 ) (G.H2)
G.h2s4 with 5 AAs PHE238@G.h2s4.1 ( 204) - THR242@G.h2s4.5 (208 ) (G.h2s4)
G.S4 with 7 AAs ALA243@G.S4.1 ( 209) - ALA249@G.S4.7 (215 ) (G.S4)
G.s4h3 with 8 AAs SER250@G.s4h3.1 ( 216) - ASN264@G.s4h3.15 (223 ) (G.s4h3) resSeq jumps
G.H3 with 18 AAs ARG265@G.H3.1 ( 224) - LEU282@G.H3.18 (241 ) (G.H3)
G.h3s5 with 3 AAs ARG283@G.h3s5.1 ( 242) - ILE285@G.h3s5.3 (244 ) (G.h3s5)
G.S5 with 7 AAs SER286@G.S5.1 ( 245) - ASN292@G.S5.7 (251 ) (G.S5)
G.s5hg with 1 AAs LYS293@G.s5hg.1 ( 252) - LYS293@G.s5hg.1 (252 ) (G.s5hg)
G.HG with 17 AAs GLN294@G.HG.1 ( 253) - ASP310@G.HG.17 (269 ) (G.HG)
G.hgh4 with 10 AAs TYR311@G.hgh4.1 ( 270) - THR320@G.hgh4.10 (279 ) (G.hgh4)
G.H4 with 27 AAs PRO321@G.H4.1 ( 280) - ARG347@G.H4.27 (306 ) (G.H4)
G.h4s6 with 11 AAs ILE348@G.h4s6.1 ( 307) - TYR358@G.h4s6.20 (317 ) (G.h4s6)
G.S6 with 5 AAs CYS359@G.S6.1 ( 318) - PHE363@G.S6.5 (322 ) (G.S6)
G.s6h5 with 5 AAs THR364@G.s6h5.1 ( 323) - ASP368@G.s6h5.5 (327 ) (G.s6h5)
G.H5 with 26 AAs THR369@G.H5.1 ( 328) - LEU394@G.H5.26 (353 ) (G.H5)
Computing distances in the interface between fragments
0
and
3
The interface is restricted to the residues within 35.0 Angstrom of each other in the reference topology.
Computing interface...done!
From 100182 potential group_1-group_2 distances, the interface was reduced to only 20307 potential contacts.
If this number is still too high (i.e. the computation is too slow) consider using a smaller interface cutoff
0%| | 0/1 [00:00<?, ?it/s]
Streaming over trajectory object nr. 0 ( 280 frames, 280 with stride 1) in chunks of 10000 frames. Now at chunk nr 0, frames so far 280
100%|███████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████████| 1/1 [00:04<00:00, 4.98s/it]
These 20 contacts capture 12.03 (~77%) of the total frequency 15.54 (over 20307 input contacts)
As orientation value, the first 31 ctcs already capture 90.0% of 15.54.
The 31-th contact has a frequency of 0.14
freq label residue idxs sum
0 1.00 D381@G.H5.13 - Q229@5.68x68 340 956 1.00
1 1.00 R385@G.H5.17 - Q229@5.68x68 344 956 2.00
2 1.00 D381@G.H5.13 - K232@5.71x71 340 959 3.00
3 0.98 Q384@G.H5.16 - I135@3.54x54 343 865 3.98
4 0.96 T350@G.h4s6.3 - R239@ICL3 309 966 4.93
5 0.85 E392@G.H5.24 - T274@6.36x36 351 976 5.79
6 0.68 Q384@G.H5.16 - Q229@5.68x68 343 956 6.47
7 0.64 H387@G.H5.19 - A134@3.53x53 346 864 7.10
8 0.57 Y391@G.H5.23 - T274@6.36x36 350 976 7.67
9 0.55 Y358@G.h4s6.20 - E237@5.76x76 317 964 8.23
10 0.49 R385@G.H5.17 - K232@5.71x71 344 959 8.71
11 0.47 L394@G.H5.26 - K270@6.32x32 353 972 9.19
12 0.47 L346@G.H4.26 - R239@ICL3 305 966 9.65
13 0.45 Y358@G.h4s6.20 - S236@5.75x75 317 963 10.11
14 0.39 Y360@G.S6.2 - S236@5.75x75 319 963 10.50
15 0.35 Q384@G.H5.16 - E225@5.64x64 343 952 10.85
16 0.34 H41@G.S1.2 - F139@34.51x51 37 869 11.19
17 0.31 Q384@G.H5.16 - T136@3.55x55 343 866 11.49
18 0.28 D139@H.hbhc.3 - K140@34.52x52 107 870 11.77
19 0.26 Y358@G.h4s6.20 - I233@5.72x72 317 960 12.03
label freq
0 Q384@G.H5.16 2.32
1 D381@G.H5.13 2.00
2 R385@G.H5.17 1.49
3 Y358@G.h4s6.20 1.26
4 T350@G.h4s6.3 0.96
5 E392@G.H5.24 0.85
6 H387@G.H5.19 0.64
7 Y391@G.H5.23 0.57
8 L394@G.H5.26 0.47
9 L346@G.H4.26 0.47
10 Y360@G.S6.2 0.39
11 H41@G.S1.2 0.34
12 D139@H.hbhc.3 0.28
label freq
0 Q229@5.68x68 2.68
1 K232@5.71x71 1.48
2 R239@ICL3 1.42
3 T274@6.36x36 1.42
4 I135@3.54x54 0.98
5 S236@5.75x75 0.85
6 A134@3.53x53 0.64
7 E237@5.76x76 0.55
8 K270@6.32x32 0.47
9 E225@5.64x64 0.35
10 F139@34.51x51 0.34
11 T136@3.55x55 0.31
12 K140@34.52x52 0.28
13 I233@5.72x72 0.26
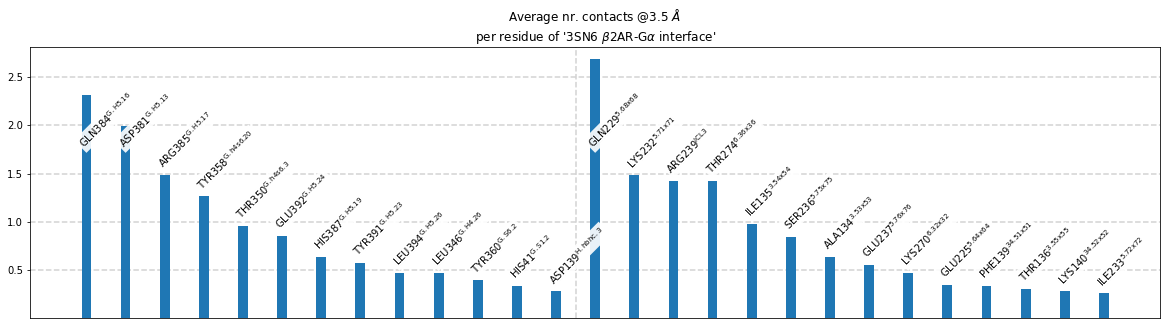
Plot each residues’s participation in the interface¶
[6]:
ifig = intf.plot_frequency_sums_as_bars(3.5, title_str = intf.name,
list_by_interface=True,
interface_vline=True);
ifig.figure.savefig("intf.svg")
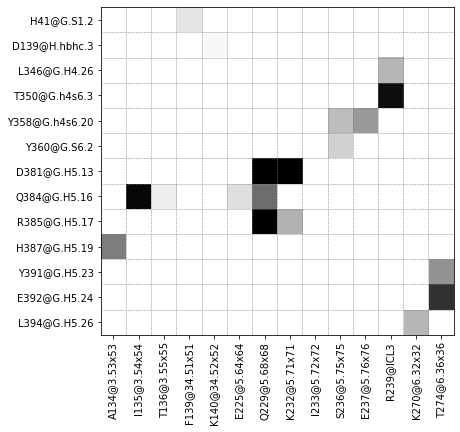
Plot contact matrix¶
[7]:
ifig, iax = intf.plot_interface_frequency_matrix(3.5, grid=True, pixelsize=.5);
ifig.savefig("matrix.svg")

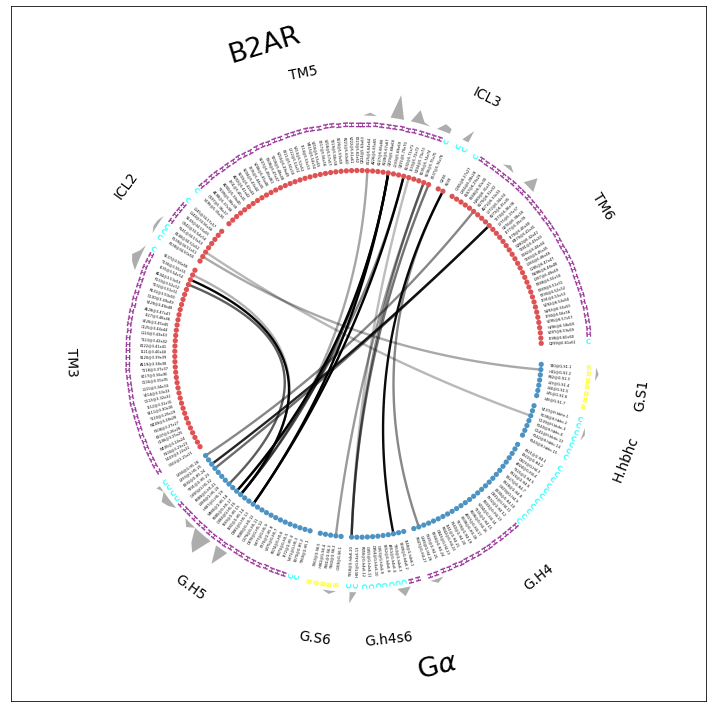
Flareplot¶
We combine a lot of information into one single flareplot:
the molecular topology with sub-fragments and consensus labels,
the secondary structure,
the individual contact-pairs
the participation of each residue in the interface.
[8]:
ifig, iax = intf.plot_freqs_as_flareplot(3.5,
fragments=fragments, fragment_names = fragment_names,
scheme="consensus_sparse", consensus_maps=[GPCR, CGN],
aura=intf.frequency_sum_per_residue_idx_dict(3.5,return_array=True),
SS=True)
ifig.figure.savefig("flare.svg")
Drawing this many dots (206 residues + 13 padding spaces) in a panel 10.0 inches wide/high
forces too small dotsizes and fontsizes. If crowding effects occur, either reduce the
number of residues or increase the panel size
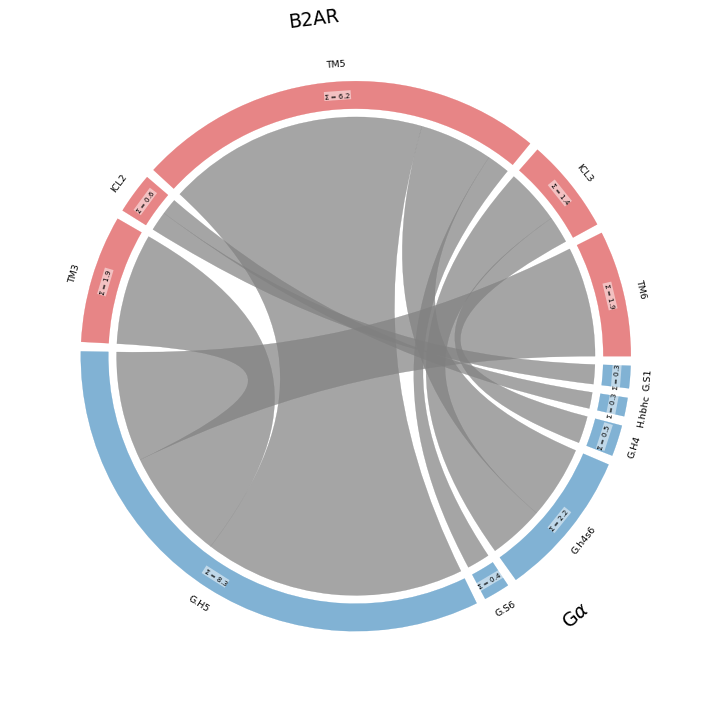
Coarse-Grained Frequencies and Flareplots¶
[9]:
ifig, iax = intf.plot_freqs_as_flareplot(3.5,
fragments=fragments, fragment_names = fragment_names,
consensus_maps=[GPCR, CGN],
coarse_grain=True,
)
intf.frequency_as_contact_matrix_CG(3.5, fragments=fragments, fragment_names = fragment_names,
consensus_labelers=[GPCR, CGN],
interface=True).round(1).replace(0,"")
[9]:
| TM3 | ICL2 | TM5 | ICL3 | TM6 | |
|---|---|---|---|---|---|
| G.S1 | 0.3 | ||||
| H.hbhc | 0.3 | ||||
| G.H4 | 0.5 | ||||
| G.h4s6 | 1.3 | 1.0 | |||
| G.S6 | 0.4 | ||||
| G.H5 | 1.9 | 4.5 | 1.9 |
Grab a representative frame¶
This frame will be used to plot the interface frequencies as a 3D heatmap (see frequency_to_bfactor below).
[10]:
repframe = intf.repframes(return_traj=True)[-1][0]
Returning frame 118 of traj nr. 0: <mdtraj.Trajectory with 280 frames, 8384 atoms, 1044 residues, and unitcells>
Save the interface as a heatmap and view externally¶
[11]:
intf.frequency_to_bfactor(3.5, pdbfile="interface_heatmap.pdb",
geom=repframe,
interface_sign=True
);
Contact frequencies stored as signed bfactor in 'interface_heatmap.pdb'
Save all mdciao objects for later reuse¶
We can save all mdciao objects to numpy .npy (pickle) files and later reload them without having to compute everything again.
[12]:
import numpy as np
np.save("GPCR.npy", GPCR)
np.save("CGN.npy",CGN)
np.save("intf.npy",intf)