FragFit
Motivation
An interactive tool for prediction, visualization and selection of missing segments in protein cryo-EM density maps.
FragFit is an interactive web service for guided modeling
of missing segments such as loops or hige regions into cryo-Electron Microscopy (cryo-EM) density maps
[1].
Segments of up to 3-35 residues length can be effectively modeled into maps of different resolution. Fragments
are taken from a pre-calculated and frequently updated database containing >1.000 million
entries (Nov 2017) extracted from structural coordinates deposited in the PDB
[2].
A fast hierarchical search algorithm (FragSearch
[3]) selects up to 100 suitable
fragments by using sequence similarity and geometrical fitting as evaluation criteria. The found fragments are then
re-ranked by their cross-correlation to the user provided cryo-EM density map. To minimize calculation time and to reduce false
positive predictions, the provided map is preprocessed.
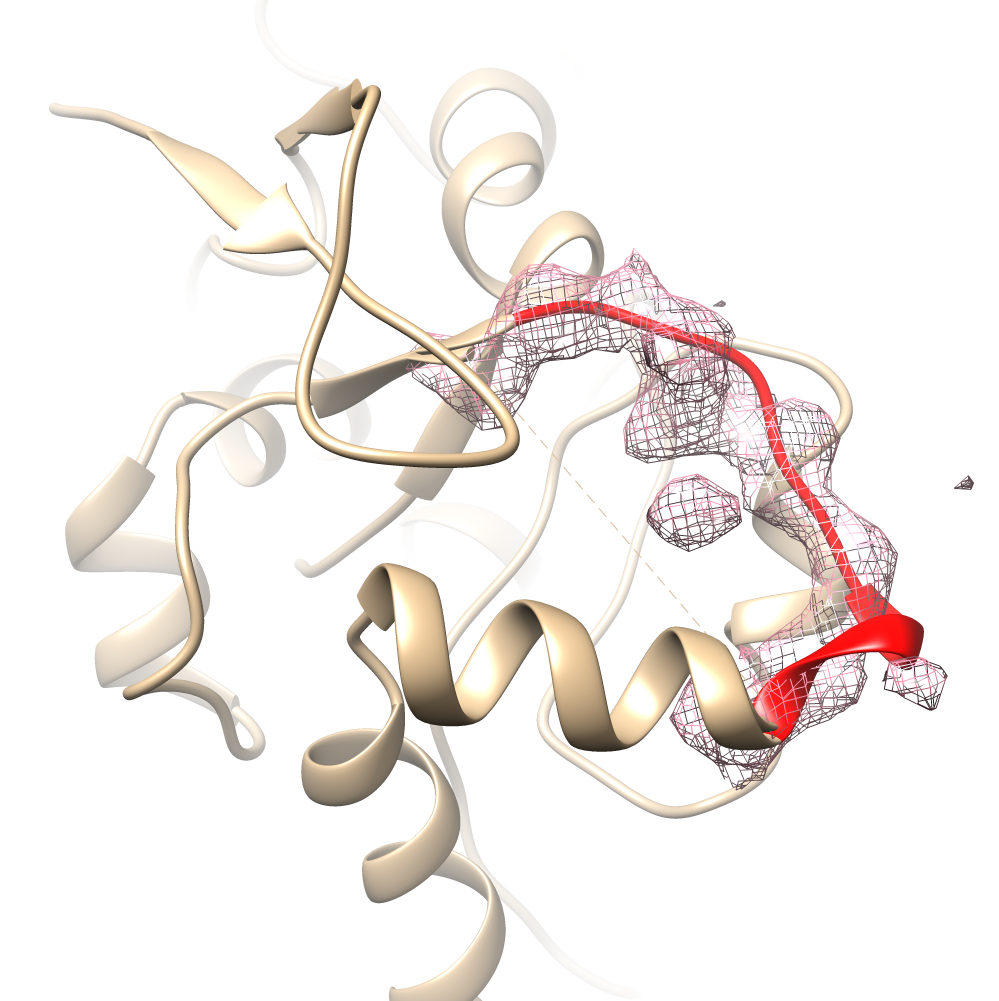
Visualization
Integration of the NGL viewer
[4], a molecular viewer based on WebGL,
allows visualization and interactive selection of a loop from a
list of 100 suitable candidates. A color code is provided to
indicate clashes or other relevant criteria to promote an
individual choice of the most suitable loop.
By inspection of the best 5 fitted fragments, prediction quality is enhanced
[1].
Results
The fragments are provided for visual inspection and can be selected from the sortable result table. A selected loop inserted into
the user provided protein can be downloaded. In addition, all 100 loop candidates can be downloaded
for further analyses.